G-spin™ Genomic DNA Extraction Kit (for Bacteria)

Description
G-spin™ Genomic DNA Extraction Mini Kit (for Bacteria) is designed to efficiently extract genomic DNA from gram-positive or gram-negative bacteria. Normally, manual methods such as alcohol precipitation are used to extract genomic DNA from bacteria, but at this time, you will feel inconvenience about process complexity and low purity. In the case of the homebrew protocol, there are so many different protocols depending on the type of bacteria, which makes it difficult to choose which method to use.
Our G-spin™ Genomic DNA Extraction Mini Kit (for Bacteria) is a dedicated product for genomic DNA extraction of various Bacteria and has been verified by many customers for a long time. An effective and milky Buffer system can stabilize the genomic DNA of bacteria and extract large size genomic DNA without shearing DNA. In addition, it is a product of Spin column type with high purity and high yield GxN technology. It provides the environment that can extract DNA most suitable for research such as PCR, Southern blot, RAPD, Sequencing.
Experimental Informations
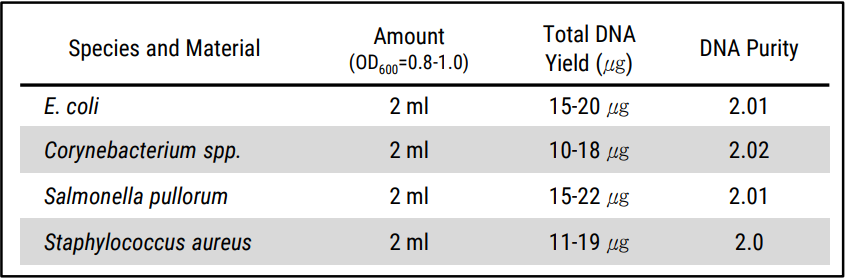
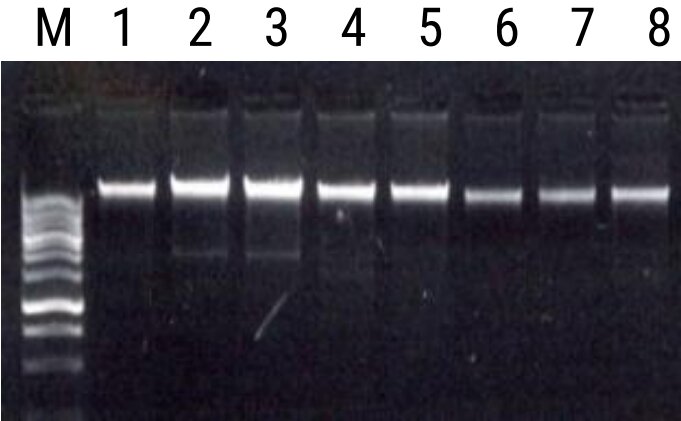
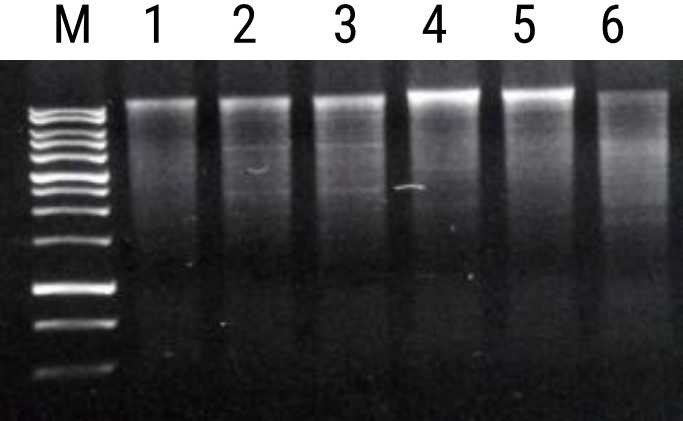

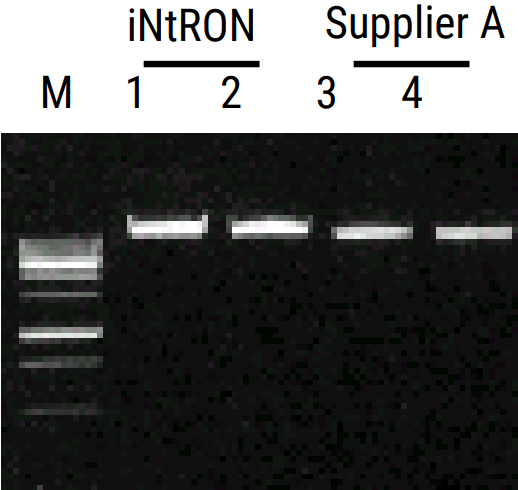
| Contents | 50 COLUMNS |
|---|---|
| Pre-Buffer | 3ml |
| G-Buffer | 20ml |
| Binding Buffer | 15ml |
| Washing Buffer A | 9ml |
| Washing Buffer B(Concentration) | 10ml |
| Elution Buffer | 20ml |
| Spin Column (Green color) & Collection tube | 50 columns |
| RNase A, lyophilized | 3 mg |
| Proteinase K, Lyophilized | 1.76mg |
| Lysozyme, Lyophilized | 20mg |
| Manual | 1 ea |
| 1 | Asian-Australas
J Anim Sci. 2017 Aug; 30(8): 1099–1104. Published online 2017 Jan 26. doi: 10.5713/ajas.16.0708 |
Effect of alcohol dehydrogenase 1C (ADH1C) genotype on vitamin A restriction and marbling in Korean native steers | Dong Qiao Peng,1,2 U Suk Jung,1 Jae Sung Lee,1,2 Won Seob Kim,1,2 Yong Ho Jo,1,2 Min Jeong Kim,1 Young Kun Oh,3 Youl Chang Baek,3 Seong Gu Hwang,4 and Hong Gu Lee1,2,* | Korea |
| 2 | Applied
Microbiology and Biotechnology July 2014, Volume 98, Issue 13, pp 6105–6113 |
Comparison of droplet digital PCR and quantitative real-time PCR for examining population dynamics of bacteria in soil | Tae Gwan KimSo-Yeon JeongKyung-Suk Cho | Korea |
| 3 | J.
Microbiol. Biotechnol. (2014), 24(6), 854–861 http://dx.doi.org/10.4014/jmb.1312.12063 |
Induction
of Immune Responses by Two Recombinant Proteins of,Brucella abortus, Outer
Membrane Proteins 2b Porin and Cu/ZnSuperoxide Dismutase, in Mouse
Model |
Kyung
Yong Sung1, Myunghwan Jung1, Min-Kyoung Shin1, Hyun-Eui Park1, Jin Ju
Lee2 , Suk Kim2, andHan Sang Yoo1,3* |
Korea |
| 4 | Mustafa, K. “Isolation and Identification of Burkholderia Mallei and Its Susceptibility to Anethum Graveolens Extracts”. ZANCO Journal of Pure and Applied Sciences, Vol. 30, no. 4, Sept. 2018, pp. 11-22, doi:10.21271/ZJPAS.30.4.2. | Isolation and Identification of Burkholderia mallei and its Susceptibility to Anethum graveolens Extracts | Khadija Khalil Mustafa | Iraq |
| 5 | 16
December 2017| https://doi.org/10.1002/biot.201700562, Cited by: 9 |
Biosynthesis of the Nylon 12 Monomer, ω‐Aminododecanoic Acid with Novel CYP153A, AlkJ, and ω‐TA Enzymes | Md. Murshidul Ahsan , Hyunwoo Jeon , Saravanan P. Nadarajan , Taeowan Chung , Hee‐Wang Yoo , Byung‐Gee Kim , Mahesh D. Patil , Hyungdon Yu | Korea |
| 6 | 01 December 2015, International Journal of Systematic and Evolutionary Microbiology 65: 4674-4681, doi: 10.1099/ijsem.0.000631 | Weissella jogaejeotgali sp. nov., isolated from jogae jeotgal, a traditional Korean fermented seafood | Se-Hui Lee1 ,5, Hye-Jin Ku1 ,5, Min-Ju Ahn1, Ji-Sang Hong1, Se Hee Lee2, Hakdong Shin3, Keun Chul Lee4, Jung-Sook Lee4, Sangryeol Ryu3, Che Ok Jeon2, Ju-Hoon Lee1 | Korea |
| 7 | Iraqi Journal of Science, Vol 63 No 6 (2022) | Detection of Efflux Pumps Gene and Relation with Antibiotics Resistance in Uropathogenic Escherichia Coli (UPEC) Isolated from Patients with Cystitis | Hussein O.M. Al-Dahmoshi | Iraq |
| 8 | Scientifc reports | Antimicrobial activity of cell free supernatants from probiotics inhibits against pathogenic bacteria isolated from fresh boar semen | Krittika Keeratikunakorn, Thotsapol Kaewchomphunuch, Kampon Kaeoket & Natharin Ngamwongsatit | Thailand |
| 9 | Journal of the Hellenic Veterinary Medical Society | Molecular identification and sequencing of Pseudomonas aeruginosa 16S rRNA gene isolated from a variety of raw cow milk samplesin Iraq | AA Saeed, KQ Mayea, SH Shalal | Iraq |
| 10 | Springer | Modified gold nanoparticle colorimetric probe-based biosensor for direct and rapid detection of Mycobacterium tuberculosis in sputum specimens | Sara Kooti, Sepide Kadivarian, Ramin Abiri, Parviz Mohajeri, Sara Atashi, Hossein Ahmadpor & Amirhooshang Alvandi | Iran |
| 11 | ELSEVIER | Molecular detection of Babesia and Theileria species/genotypes in sheep and ixodid ticks in Erzurum, Northeastern Turkey: First report of Babesia canis in sheep | Ridvan Kirman, Esin Guven | Turkey |
| Product | Cat.No | Capacity | inquire | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| i-Taq™ DNA Polymerase |
|
|||||||||
| i-StarTaq™ DNA Polymerase Best |
|
|||||||||
| Maxime™ PCR PreMix (i-StarMAXⅡ) |
|
|||||||||
| SiZer™-1,000 DNA Marker Solution Best |
|
|||||||||
| SiZer™-1,000 plus DNA Marker Solution |
|
|||||||||
| RedSafe™ Nucleic Acid Staining Solution Best |
|
|||||||||